Research
Graduate Research
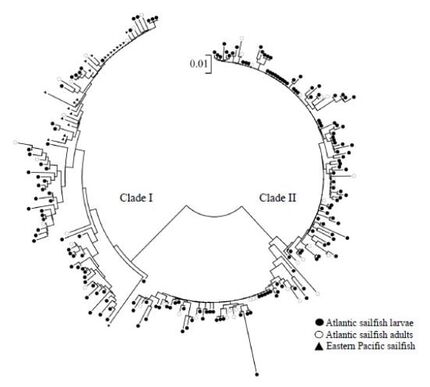
A clear understanding of the genetic population structure and genetic diversity of threatened species is critical when devising management practices for conservation. My career in conservation genomics began during my Master’s degree in wildlife and fisheries science, where I used genetic techniques to assess the extent of population structure in sailfish, a marine fish species of important conservation concern. I used a combination of microsatellite loci and mitochondrial sequence data to assess the levels of genetic diversity within and genetic structure between two Atlantic (eastern and western) and one Pacific (eastern) populations of sailfish. Most notably, I detected modest yet significant population structure between eastern and western populations of Atlantic sailfish as resolved using our panel microsatellite loci. To our knowledge this is the first reported detection of such structure between Atlantic populations of this species. I also used our mitochondrial sequence data to investigate the historical demography of both Atlantic and Pacific populations of sailfish and invoke the “Mitchell-Perrin model of unidirectional movement” to describe the predominant pattern of gene flow from the Indo-Pacific into the Atlantic as facilitated by the westward flowing Agulhas current.
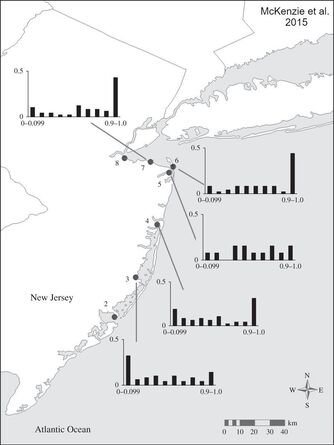
In my PhD thesis, I integrated field work, laboratory experiments, and genetic techniques to investigate the strength of selection and role of reproductive isolation in maintaining a hybrid zone between subspecies of a model teleost, the killifish (Fundulus heteroclitus), whose range coincides with a steep latitudinal thermal gradient along North America’s Atlantic coast. Hybrid zones are areas where genetically distinct taxa meet and interbreed and provide natural laboratories for the study of the processes involved in adaptive divergence and the evolution of reproductive isolation. I demonstrated that local adaptation to differing thermal conditions is associated with partial reproductive isolation between the two subspecies of killifish. Given that hybridization between native and non-native species of fish is a significant conservation concern in many jurisdictions, increasing our understanding of the relative roles of local adaptation and reproductive isolation in maintaining species boundaries has significant conservation implications.
Postdoctoral Research
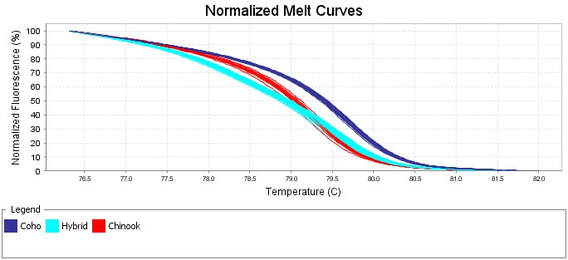
My first postdoctoral position was as a Natural Sciences and Engineering Research Council of Canada (NSERC) Visiting Fellow at Fisheries and Oceans Canada. My research during this time reinforced my interest in hybridization among closely related taxa as both a creative and destructive force. I used whole animal behavioural experiments as well as next-generation sequencing (RNAsequencing) to examine the degree of reproductive isolation between four species of Pacific salmon as well as the transcriptomic effects of hybridization between these species. I took full advantage of the opportunity to become proficient with several bioinformatic pipelines including those for transcriptome assembly and annotation, differential gene expression, gene ontology analyses, SNP calling, and allele specific expression.
I am currently a postdoctoral researcher at the University of British Columbia where we are using rainbow trout as a model to investigate the intersection between genotype, phenotype, and climate change. This project is allowing me the opportunity to work on a contemporary problem in conservation biology in the current context of rapid global climate change. We have collected physiological phenotypes (critical thermal maximum and the ability to tolerate low oxygen stress) for hundreds of individuals belonging to several strains of rainbow trout and subsequently using genomic data (genotyping-by-sequencing) to make genotype-phenotype associations (via genome-wide association studies) with the long-term goal of identifying a strain of rainbow trout best-suited to tolerate the changing environment we live in.
Future Research Plans
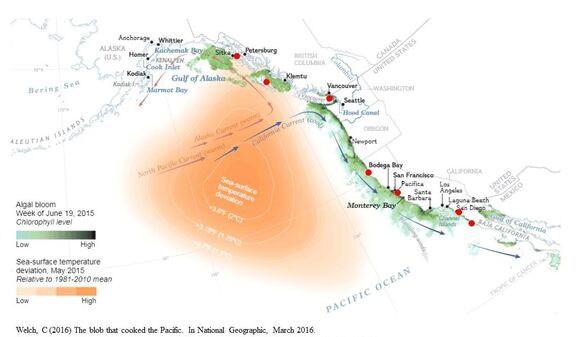
I am preparing a proposal for submission to NSERC's Alliance Grant Program to fund a project entitled “Climate change and genetic diversity: Benefits and threats for British Columbia's endangered pinto abalone (Haliotis kamtschatkana)” . I will use a combination of next generation sequencing techniques (whole genome sequencing, RNAsequencing, and genotyping-by-sequencing) to assess the species status of British Columbia’s pinto abalone, compare genetic diversity among populations throughout this species’ range, and define the location of the hybrid zone connecting pinto and threaded types. Laboratory experiments will investigate how pinto and threaded abalone differentially regulate their responses to environmental extremes. As the range of the pinto abalone encompasses three countries, this project will be a large-scale international effort. I have previously obtained commitment from collaborators within government (NOAA, USA; Fisheries and Oceans Canada; California Department of Fish and Wildlife), academia (University of California Santa Cruz, USA; Ensenada Center for Scientific Research and Higher Education, Baja, California, Mexico; University of California Davis, USA), and non-profit organizations (Puget Sound Restoration Fund, USA; Sitka Sound Science Centre; Paua Marine Research Group) throughout the species range from Canada, through the United States, and into northern Mexico.
